But this post talks about a different idea, and it is awesome. Now that they have all this geotagged pictures and they know they are in a particular neighborhood or country (WOEID) they are creating shapefiles from these areas based on the coordinates of the different pictures! Maybe is easier to explain it with a picture:

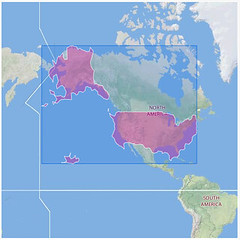
The one in the left is a London polygon and the right for United States. They were created by aggregating all the coordinates of the different pictures found in London and in the US.
Again. They take all the coordinates, POINTs, in the DB for a certain WOEID and with them they generate a polygon. If they have enough data it looks more or less ok. They have a good discussion about the quality of the polygons and the threshold set to generate them. I like very much the introduction they make to Alpha shapes and the links they provide.
Additionally they also provide the source code of the tool they use to process the data. It is called Clustr.
Now, lets apply this concept to Biodiversity, if you haven't already figure it out. Think of their geotagged pictures as PRIMARY DATA, the WOE as SCIENTIFIC NAMES and the polygons you get out of them as DERIVED SPECIES DISTRIBUTION polygons.
I have been investigating this for a long time already, specially trough the Biodiversity Atlas project that is now stopped, but hopefully will start soon. We can take their source code and apply it to GBIF data and generate "derived, unchecked and uncompleted" species distributions based on GBIF data on a massive way! And the same they try in Flickr, the more primary data we get into the system the better the distributions will start looking like.
But there is another idea... why dont ask Flickr to not only process their polygons based on WOE Ids but also on tags? So if we tag pictures in Flickr with scientific names, or better GUIDs, they can then try to generate by themselves the distributions.
I particularly dont see Flickr as the best place to handle species distributions discussions, and will, hopefully, try to convince at least one big biodiversity project to let me try this way. Most of you can probably imagine the incredible API we can create once we have a lot of species distributions accessible in such a system. I will write another post about it soon, but think of:
- Which species could live in my garden?
- Which species habitats this new road will cross?
- Where should I create a new Protected Area to preserve as much biodiversity as possible?
- What species could I find in the track I will hike this weekend?
So maybe instead of so much niche modeling projects we should start thinking on how to manage the vast amount of primary data (and other sources) we have already and how to curate it and complete it. I dream of a scientific community joined together to create a complete information system to know where species are. Imagine like a Wikipedia but for Species Distributions.




4 comments:
Well, this will hopefully become reality one day. But there are many problems with biodiversity occurrence data that Flickr doesnt deal with. They use reverse geocoding to obtain the WOEID, so they never get really bad outliers. But our coordinates and taxon names are completely decoupled. So we have a lot of wrong points because of various reasons: synonyms and lack of species GUIDs, many wrong identifications, leopards sighted in European zoos ("basis of record"). Also some people think of distribution probably as the "native" distribution and would not want to count invasives.
The bottom line for me is that there really needs to be a quality indication for occurrences/points, not all of them are equally reliable. And to detect bad records you would want species distibutions - back to where we started.
Well, Flickr also have bad outliners. Specially at the neighborhood level. For this they can not directly reverse geocode as there is no official neighborhood shapes. They are looking for discussion between users to delimitate them. So they also have to to explain that this data will never be correct, it will be permanently being edited and what you see is more like a zeitgeist of a place.
They have also points that are totally wrong. Flickr automatically reverse geocode, but you can edit it and change it, and they have to decide if the isolate it point, Madrid in the Antarctic, make sense or not for inclusion. The alpha shape algorithm can help discard some of this wrong data.
Taxonomy of course is a big problem. But I was wondering some days ago some idea. Has synonymy a huge impact in distributions? The worst scenario is that a map is not showing all data you have, and it will never actually as this is a matter of taxonomic opinion. Again, the best you can do is show the zeitgeist of a taxonomic name.
Wrong identifications of course will always be an issue, but hopefully scaling the system and allowing for "experts" to comments and delimit areas could help on that.
Regarding the basis of records this must be model in the system. And also the status. But I would do something simpler than TDWG (http://www.nhm.ac.uk/hosted_sites/tdwg/poss_standard.html).
I agree on your bottom line. But we are a a little bit luckier than Flickr on this. We have different sources f species distributions, not only primary data, all this together, can provide a system where the data can be continuously improved and checked with different sources of information.
Guys,
Nice idea and very interesting discussion. In my humble opinion I think that a system like Life Mapper is theoretically better better than what you have in mind because it is using environmental data to predict and model species distributions (which should be superior predictor of species ditribution than pure geographical space). Assuming of course that whatever data you have available for your Flickr distributions was also be available to LifeMapper. How is your idea better than a LifeMapper model for each species.
Would it not be better to use the LifeMapper models to answer the questions - what would occur in my garden, what species habitat would a road cross etc.
The problem with LifeMapper and the Flickr approach is always going to be dodgy data (though admittedly an outlier will affect LifeMapper models more than the Flickr distribution).
One possibility of improving your suggestion is to incorporate some form of probability into the FLickr polygons (yikes - how do we do that within a polygon) where those neighborhoods where only one point occurs has lower probability of species occuring there than those with 2 or more etc.
I look forward to some robust debate on this matter
Cheers
Paul
Hi Paul,
I think what is missing in this post is the context of why we are discussing these techniques about deriving polygons from all kinds of point data.
In a very brief summary, we are discussing an overarching system that allows for indexing and merging of datasources providing species distribution information, and also authoring of the distribution using grid drawing, polygon drawing etc. The idea is that if you have as many sources as possible (including those derived from occurrence data - see other blog posts on this but occurrence data as points have severe performance issues, so we always need to polygon or grid the points), and keeping them as separate "layers", a system could be built that allows for a whole community to quickly annotate / rank / author modifications to layers and produce something that resembles a "real" species distribution (temporal component is included also). Our thinking is that by incorporating enough sources, the dodgy data would fall out the bottom as outliers, so basically probability based like you suggest but across multiple sources. Datasource type (basis of record) would be another requirement to indicate if the polygon was derived or considered authoritative. The output of such a system would both provide distribution data and a means to help validate the point data.
So the output of environmental layers could be another source to consider, but does not fit the parameters quite so well, since it is producing distribution of where the species *could* potentially live, rather than known to live - both systems would complement each other though being used to help validate the others outputs, and perhaps this could be used as a model input the the niche models? Here is the best known distribution (not only some points) of SpeciesX today - run a 2050 climate model on it - although I am not sure if niche modeling systems can take a polygon based input.
Cheers,
Tim
Post a Comment